CCA
Canonical Correlation Analysis was first proposed by Hotelling (1936) 🎓. It can be conceived as the multivariate extension of Pearson's product-moment correlation, as the bilinear version of Whitening or as the standardized version of MCA; if MCA maximizes the cross-covariance of two data set, CCA maximizes their correlation. Like the MCA, the CCA corresponds to the situation $m=2$ (two datasets) and $k=1$ (one observation).
As per MCA, Let $X$ and $Y$ be two $n_x⋅t$ and $n_y⋅t$ data matrices, where $n_x$ and $n_y$ are the number of variables in $X$ and $Y$, respectively and $t$ the number of samples. We assume here too that the samples in $X$ and $Y$ are synchronized. Let $C_{x}$ be the $n_x⋅n_x$ covariance matrix of $X$ and $C_{y}$ be the $n_y⋅n_y$ covariance matrix of $Y$. Let also $C_{xy}=\frac{1}{t}X^HY$ be the $n_x⋅n_y$ cross-covariance matrix. CCA seeks two matrices $F_x$ and $F_y$ such that
$\left \{ \begin{array}{rl}F_x^HC_xF_x=I\\F_y^HC_yF_y=I\\F_x^HC_{xy}F_y=Λ \end{array} \right.$, $\hspace{1cm}$ [cca.1]
where $Λ$ is a $n⋅n$ diagonal matrix, with $n=min(n_x, n_y)$. The first components (rows) of $F_x^{H}X$ and $F_y^{H}Y$ hold the linear combination of $X$ and $Y$ with maximal correlation, the second the linear combination with maximal residual correlation and so on, subject to constraint [cca.1].
In matrix form this can be written such as
$\begin{pmatrix} F_x^H & 0\\0 & F_y^H\end{pmatrix} \begin{pmatrix} C_x & C_{xy}\\C_{xy}^H & C_y\end{pmatrix} \begin{pmatrix} F_x & 0\\0 & F_y\end{pmatrix} = \begin{pmatrix} I & Λ\\Λ & I\end{pmatrix}$. $\hspace{1cm}$ [cca.2]
If $n_x=n_y$, from [cca.1] it follows
$\left \{ \begin{array}{rl}F_x^{-H}F_x^{-1}=C_x\\F_y^{-H}F_y^{-1}=C_y\\F_x^{-H}F_y^{-1}=C_{xy} \end{array} \right.$, $\hspace{1cm}$ [cca.3]
that is, CCA is a special kind of full-rank factorization of $C_x$ and $C_y$.
Equating the left hand sides of the first two expressions in [cca.1] and setting $E_x=B_xB_y^{-H}$, $E_y=B_yB_x^{-H}$, it follows
$\left \{ \begin{array}{rl}C_x=E_xC_yE_x^H\\C_y=E_yC_xE_y^H \end{array} \right.$, $\hspace{1cm}$ [cca.4]
that is, CCA defines a linear transormation linking the covariance matrices of two data sets.
Since it maximizes the correlation, differently from MCA, CCA is not sensitive to the amplitude of the two processes. This should be kept in mind, as the correlation of two signals may be high even if the amplitude of one of the two signal is very low, say, at the noise level. In such a case the resulting correlation is meaningless.
The accumulated regularized eigenvalues (arev) for the CCA are defined as
$σ_j=\sum_{i=1}^j{σ_i}$, for $j=[1 \ldots n]$, $\hspace{1cm}$ [cca.5]
where $σ_i$ is given by
$σ_p=\frac{\sum_{i=1}^pλ_i}{σ_{TOT}}$ $\hspace{1cm}$ [cca.6]
and $λ_i$ are the singular values in $Λ$ of [cca.1].
For setting the subspace dimension $p$ manually, set the eVar optional keyword argument of the CCA constructors either to an integer or to a real number, this latter establishing $p$ in conjunction with argument eVarMeth using the arev vector (see subspace dimension). By default, eVar is set to 0.999.
Solution
If $n_x=n_y$ the solutions to the CCA $B_x$ and $B_y$ are given by the eigenvector matrix of (non-symmetric) matrices $C_x^{-1}C_{xy}C_y^{-1}C_{yx}$ and $C_y^{-1}C_{yx}C_x^{-1}C_{xy}$, respecively.
A numerically preferable solution, accommodating also the case $n_x≠n_y$, is the following two-step procedure, which is the one here implemented:
- get some whitening matrices $\hspace{0.1cm}(W_x, W_y)\hspace{0.1cm}$ such that $\hspace{0.1cm}W_x^HC_xW_x=I\hspace{0.1cm}$ and $\hspace{0.1cm}W_y^HC_yW_y=I$
- do $\hspace{0.1cm}\textrm{SVD}(W_x^HC_{xy}W_y)=U_xΛU_y^{H}$
The solutions are $\hspace{0.1cm}B_x=W_xU_x\hspace{0.1cm}$ and $\hspace{0.1cm}B_y=W_yU_y\hspace{0.1cm}$.
Constructors
Three constructors are available (see here below). The constructed LinearFilter object holding the CCA will have fields:
.F[1]: matrix $\widetilde{B}_x=[b_{x1} \ldots b_{xp}]$ with the columns holding the first $p$ vectors in $B_x$.
.F[2]: matrix $\widetilde{B}_y=[b_{y1} \ldots b_{yp}]$ with the columns holding the first $p$ vectors in $B_y$.
.iF[1]: the left-inverse of .F[1]
.iF[2]: the left-inverse of .F[2]
.D: the leading $p⋅p$ block of $Λ$, i.e., the correlation values associated to .F in diagonal form.
.eVar: the explained variance for the chosen value of $p$, defined in [cca.6].
.ev: the vector diag(Λ) holding all $n$ correlation values.
.arev: the accumulated regularized eigenvalues, defined in [cca.5].
Diagonalizations.cca — Function
(1)
function cca(Cx :: SorH, Cy :: SorH, Cxy :: Mat;
eVarCx :: TeVaro=○,
eVarCy :: TeVaro=○,
eVar :: TeVaro=○,
eVarMeth :: Function=searchsortedfirst,
simple :: Bool=false)
(2)
function cca(X::Mat, Y::Mat;
covEst :: StatsBase.CovarianceEstimator=SCM,
dims :: Into = ○,
meanX :: Tmean = 0,
meanY :: Tmean = 0,
wX :: Tw = ○,
wY :: Tw = ○,
wXY :: Tw = ○,
eVarCx :: TeVaro = ○,
eVarCy :: TeVaro = ○,
eVar :: TeVaro = ○,
eVarMeth :: Function = searchsortedfirst,
simple :: Bool = false)
(3)
function cca(𝐗::VecMat, 𝐘::VecMat;
covEst :: StatsBase.CovarianceEstimator=SCM,
dims :: Into = ○,
meanX :: Into = 0,
meanY :: Into = 0,
eVarCx :: TeVaro = ○,
eVarCy :: TeVaro = ○,
eVar :: TeVaro = ○,
eVarMeth :: Function = searchsortedfirst,
simple :: Bool = false,
metric :: Metric = Euclidean,
wCx :: Vector = [],
wCy :: Vector = [],
✓w :: Bool = true,
initCx :: SorHo = nothing,
initCy :: SorHo = nothing,
tol :: Real = 0.,
verbose :: Bool = false)Return a LinearFilter object:
(1) Canonical correlation analysis with, as input, real or complex:
- covariance matrix
Cxof dimension $n_x⋅n_x$, - covariance matrix
Cyof dimension $n_y⋅n_y$ and - cross-covariance matrix
Cxyof dimension $n_x⋅n_y$.
Cx and Cy must be flagged as Symmetric, if real or Hermitian, if real or complex, see data input. Instead Cxy is a generic Matrix object since it is not symmetric/Hermitian, left alone square.
eVarCx, eVarCy, eVar and eVarMeth are keyword optional arguments for defining the subspace dimension $p$. Particularly:
eVarCxandeVarCyare used to determine a subspace dimension in the whitening ofCxandCy, i.e., in the first step of the two-step procedure descrived above in the solution section.eVaris used to determine the final subspace dimension using the.arevvector given by Eq. [cca.5] here above.eVarMethapplies toeVarCx,eVarCyandeVar.
The default values are:
eVarCx=0.999eVarCy=0.999eVar=0.999eVarMeth=searchsortedfirst
If simple is set to true, $p$ is set equal to $min(n_x, n_y)$ and only the fields .F and .iF are written in the constructed object. This option is provided for low-level work when you don't need to define a subspace dimension or you want to define it by your own methods.
(2) Canonical correlation analysis with real or complex data matrices X and Y as input.
covEst, dims, meanX, meanY, wX, wY and wXY are optional keyword arguments to regulate the estimation of the covariance matrices of X and Y and their cross-covariance matrix $C_{xy}$. Particularly (See covariance matrix estimations),
meanXis themeanargument for data matrixX.meanYis themeanargument for data matrixY.wXis thewargument for estimating a weighted covariance matrix $C_x$.wYis thewargument for estimating a weighted covariance matrix $C_y$.wXYis thewargument for estimating a weighted cross-covariance matrix $C_{XY}$.covEstapplies only to the estimation of $C_x$ and $C_y$.
Once $C_x$, $C_y$ and $C_{xy}$ estimated, method (1) is invoked with optional keyword arguments eVar, eVarCx, eVarCy, eVarMeth and simple. See method (1) for details.
(3) Canonical correlation analysis with two vectors of real or complex data matrices 𝐗 and 𝐘 as input. 𝐗 and 𝐘 must hold the same number of matrices and corresponding pairs of matrices therein must comprise the same number of samples.
covEst, dims, meanX and meanY are optional keyword arguments to regulate the estimation of the covariance matrices of all matrices in 𝐗 and 𝐘 and the cross-covariance matrices for all pairs of their corresponding data matrices. See method (2) and covariance matrix estimations.
A mean of these covariance matrices is computed. For the cross-covariance matrices the arithmetic mean is used. For the covariance matrices of 𝐗 and 𝐘, optional keywords arguments metric, wCx, wCy, ✓w, initCx, initCy, tol and verbose are used to allow non-Euclidean mean estimations. Particularly (see mean covariance matrix estimations),
wCxare the weights for the covariance matrices of𝐗,wCyare the weights for the covariance matrices of𝐘,initCxis the initialization for the mean of the covariance matrices of𝐗,initCyis the initialization for the mean of the covariance matrices of𝐘.
By default, the arithmetic mean is computed.
Once the mean covariance and cross-covariance matrices are estimated, method (1) is invoked with optional keyword arguments eVarCx, eVarCy, eVar, eVarMeth and simple. See method (1) for details.
See also: Whitening, MCA, gCCA, mAJD.
Examples:
using Diagonalizations, LinearAlgebra, PosDefManifold, Test
# Method (1) real
n, t=10, 100
X=genDataMatrix(n, t)
Y=genDataMatrix(n, t)
Cx=Symmetric((X*X')/t)
Cy=Symmetric((Y*Y')/t)
Cxy=(X*Y')/t
cC=cca(Cx, Cy, Cxy, simple=true)
@test cC.F[1]'*Cx*cC.F[1]≈I
@test cC.F[2]'*Cy*cC.F[2]≈I
D=cC.F[1]'*Cxy*cC.F[2]
@test norm(D-Diagonal(D))+1. ≈ 1.
# Method (1) complex
Xc=genDataMatrix(ComplexF64, n, t)
Yc=genDataMatrix(ComplexF64, n, t)
Cxc=Hermitian((Xc*Xc')/t)
Cyc=Hermitian((Yc*Yc')/t)
Cxyc=(Xc*Yc')/t
cCc=cca(Cxc, Cyc, Cxyc, simple=true)
@test cCc.F[1]'*Cxc*cCc.F[1]≈I
@test cCc.F[2]'*Cyc*cCc.F[2]≈I
Dc=cCc.F[1]'*Cxyc*cCc.F[2]
@test norm(Dc-Diagonal(Dc))+1. ≈ 1.
# Method (2) real
cXY=cca(X, Y, simple=true)
@test cXY.F[1]'*Cx*cXY.F[1]≈I
@test cXY.F[2]'*Cy*cXY.F[2]≈I
D=cXY.F[1]'*Cxy*cXY.F[2]
@test norm(D-Diagonal(D))+1. ≈ 1.
@test cXY==cC
# Method (2) complex
cXYc=cca(Xc, Yc, simple=true)
@test cXYc.F[1]'*Cxc*cXYc.F[1]≈I
@test cXYc.F[2]'*Cyc*cXYc.F[2]≈I
Dc=cXYc.F[1]'*Cxyc*cXYc.F[2]
@test norm(Dc-Diagonal(Dc))+1. ≈ 1.
@test cXYc==cCc
# Method (3) real
# canonical correlation analysis of the average covariance and cross-covariance
k=10
Xset=[genDataMatrix(n, t) for i=1:k]
Yset=[genDataMatrix(n, t) for i=1:k]
c=cca(Xset, Yset)
# ... selecting subspace dimension allowing an explained variance = 0.9
c=cca(Xset, Yset; eVar=0.9)
# ... subtracting the mean from the matrices in Xset and Yset
c=cca(Xset, Yset; meanX=nothing, meanY=nothing, eVar=0.9)
# cca on the average of the covariance and cross-covariance matrices
# computed along dims 1
c=cca(Xset, Yset; dims=1, eVar=0.9)
# name of the filter
c.name
using Plots
# plot regularized accumulated eigenvalues
plot(c.arev)
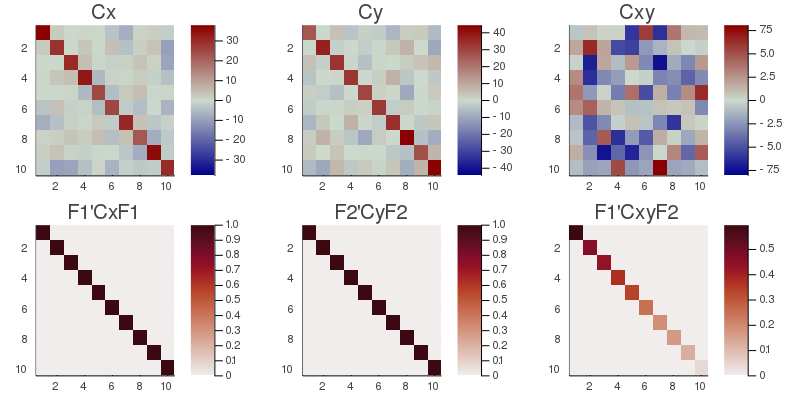
# plot the original covariance and cross-covariance matrices
# and their transformed counterpart
CxyMax=maximum(abs.(Cxy));
h1 = heatmap(Cxy, clim=(-CxyMax, CxyMax), title="Cxy", yflip=true, c=:bluesreds);
D=cC.F[1]'*Cxy*cC.F[2];
Dmax=maximum(abs.(D));
h2 = heatmap(D, clim=(0, Dmax), title="F1'CxyF2", yflip=true, c=:amp);
CxMax=maximum(abs.(Cx));
h3 = heatmap(Cx, clim=(-CxMax, CxMax), title="Cx", yflip=true, c=:bluesreds);
h4 = heatmap(cC.F[1]'*Cx*cC.F[1], clim=(0, 1), title="F1'CxF1", yflip=true, c=:amp);
CyMax=maximum(abs.(Cy));
h5 = heatmap(Cy, clim=(-CyMax, CyMax), title="Cy", yflip=true, c=:bluesreds);
h6 = heatmap(cC.F[2]'*Cy*cC.F[2], clim=(0, 1), title="F2'CyF2", yflip=true, c=:amp);
📈=plot(h3, h5, h1, h4, h6, h2, size=(800,400))
# savefig(📈, homedir()*"\Documents\Code\julia\Diagonalizations\docs\src\assets\FigCCA.png")
# Method (3) complex
# canonical correlation analysis of the average covariance and cross-covariance
k=10
Xcset=[genDataMatrix(ComplexF64, n, t) for i=1:k]
Ycset=[genDataMatrix(ComplexF64, n, t) for i=1:k]
cc=cca(Xcset, Ycset)
# ... selecting subspace dimension allowing an explained variance = 0.9
cc=cca(Xcset, Ycset; eVar=0.9)
# ... subtracting the mean from the matrices in Xset and Yset
cc=cca(Xcset, Ycset; meanX=nothing, meanY=nothing, eVar=0.9)
# cca on the average of the covariance and cross-covariance matrices
# computed along dims 1
cc=cca(Xcset, Ycset; dims=1, eVar=0.9)
# name of the filter
cc.name